Horn of Africa
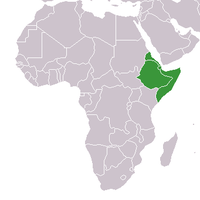
The Horn of Africa (alternatively Northeast Africa, and sometimes Somali Peninsula) is a peninsula in East Africa that sticks out into the Somali Sea and Guardafui Channel, and is along the southern side of the Gulf of Aden.
Pan-Hornism
changeSome people said that its a good idea for Horn Africans to help each other with making money and politics. Or in general This aim is sometimes called Pan-Hornism.[1][2][3]
Borders
changeThere are four countries in the Horn of Africa. They are Eritrea, Ethiopia, Somalia and Djibouti. The northernmost areas of the Horn are Ras Kasar in the west, settled by the Rashaida tribe, and Alula in the east, settled by Dishiishe. The easternmost area is Ras Hafun, settled by Somalis, and the southernmost in Ras Kamboni, settled by Somalis.[4]
Northeast Africa
changeNortheast Africa, or Northeastern Africa or Northern East Africa as it was known in the past, is a geographic regional term used to refer to the countries of Africa in and around the Red Sea. The region is between North Africa and East Africa, and mainly encompasses the Horn of Africa (Ethiopia, Eritrea, Somalia, Djibouti) and the Sudans (Sudan and South Sudan) as well as Egypt. Sometimes, its borders are stretched to either include Kenya, or Chad and Libya. Humans have lived in the area for a long time. Many fossil from the early hominids to modern human have been found here. It is one of the most genetically and linguistically diverse regions of the world, being the home to many civilizations and located on an important trade route that connects multiple continents.[5][6][7][8][9]
History
changeThe Horn of Africa became famous for taking longer to be colonized than other African countries. Those countries which fought back were Ethiopia, led by Menelik II, and the Daraawiish State, led by Ismail Mire.[10] Eritrea was part of Ethiopia, but fought the Eritrean War of Independence from 1961 to 1991.[11]
Numbers
changeHorn of Africa can also mean the greater region containing the countries of Eritrea, Djibouti, Ethiopia and Somalia. The greater region covers about 2,000,000 km2 (or 772,200 square miles) and is inhabited by about 94.2 million people (Ethiopia: 79 million, Somalia: 10 million, Eritrea: 4.5 million, and Djibouti: 0.7 million).
Genetics
changeUniparental lineages
changeCushitic ethnic groups have a diverse set of uniparental lineages. Nevertheless, certain commonalities can be observed.
Paternally, E-M35 (also known as E1b1b1, formerly E3b1) forms an important lineage in many Cushitic populations, other important paternal lineages in Cushitic populations include J-M267 (also known as J1), A-M13 (A1b1b2b, formerly A3b2), and T-M70 (T1a, formerly K2).[12][13][14][15][16][17]
Many Cushitic populations can be paternally traced back to having ethnic origins in the Nile Valley (Egypt and Northern Sudan) through haplogroup E-M78 and the Red Sea region of the Horn of Africa through haplogroup E-V1515 This coincides with anthropological and linguistic hypotheses placing the ethnogensis of the Cushitic language family in the aforementioned regions.[18]
Maternally, Cushitic populations are more diverse, yet share certain lineages in common such as various East African-origin Macro-haplogroup L lineages (various L0, L1, L5, L2, L6, L4, L3 lineages) and North African and/or Middle Eastern-origin M1 and Macro-haplogroup N (in particular N subclades N1a, N1b, R0a, HV1b1, I, K1a, U3a, and U6a).[19][20][21]
Autosomal ancestry
changeCushitic populations have existed at the cross-roads of Africa and Eurasia since the Stone Age, with the Nile acting as a corridor between sub-Saharan Africa and Levant and North Africa thus Cushitic populations have multiple origins, like most other populations, that have become idiosyncratic of Cushitic populations. Cushitic populations tend to combine in their ancestries both, genetic components indigenous to East Africa, and non-African components of West Asian origin. According to an autosomal DNA study by Hodgson et al. (2014), the Afro-Asiatic languages were likely spread across Africa and the Near East by an ancestral population(s) carrying a newly identified non-African genetic component, which the researchers dub the “Ethio-Somali” or “Semitic-Cushitic” in another study. This Ethio-Somali component is today most common among Cushitic and Ethiosemitic populations in the Horn of Africa and reaches a frequency peak among ethnic Somalis, representing the majority of their ancestry. The Ethio-Somali component is most closely related to the Maghrebi non-African genetic component, and is believed to have diverged from all other non-African ancestries at least 23,000 years ago. On this basis, the researchers suggest that the original Ethio-Somali carrying population(s) probably arrived in the pre-agricultural period from the Near East, having crossed over into northeastern Africa via the Sinai Peninsula. The population then likely split into two branches, with one group heading westward toward the Maghreb and the other moving south into the Horn.[22] Ancient DNA analysis indicates that this foundational ancestry in the Horn region is akin to that of the Neolithic farmers of the southern Levant.[23]
According to Hodgson et al. (2014), both the African ancestry (Ethiopic) and the non-African ancestry (Ethio-Somali) in Cushitic speaking populations is significantly differentiated from all neighboring African and non-African ancestries in East Africa, North Africa, the Levant and Arabia. The genetic ancestry of Cushitic and Semitic speaking populations in the Horn of Africa represents ancestries (Ethiopic and Ethio-Somali) not found outside of Cushitic and Semitic speaking HOA populations in any significance. Therefore, both ancestries are distinct, unique to, and considered the signature autosomal genetic ancestry of Cushitic and Semitic speaking HOA populations.
Hodgson et al. states:
”The African Ethiopic ancestry is tightly restricted to HOA populations and likely represents an autochthonous HOA population. The non-African ancestry in the HOA, which is primarily attributed to a novel Ethio-Somali inferred ancestry component, is significantly differentiated from all neighboring non-African ancestries in North Africa, the Levant, and Arabia.”[22]
According to Hodgson et al. (2014), the non-African ancestry (Ethio-Somali) in the Cushitic speaking populations is distinct and unique to Cushitic and Semitic speaking HOA populations. Hodgson et al. states:
”We find that most of the non-African ancestry in the HOA can be assigned to a distinct non-African origin Ethio-Somali ancestry component, which is found at its highest frequencies in Cushitic and Semitic speaking HOA populations.”[22]
When calculated the genetic distance (FST) between Ethiosemitic-speaking and Cushitic-speaking Ethiopians, and populations of the Levant, North Africa, and the Arabian Peninsula using two approaches: (1) the whole genome and (2) only the non-African component — in the whole-genome analysis, Ethiopian Semitic and Cushitic populations appear to be closest to the Yemeni; when only the non-African component is used, they are closer to the Egyptians and populations inhabiting the Levant.[24] However this is due to the fact that the similarity is because of the Ethiopian contribution to the Yemeni gene pool. Pagani et al. (2012) states:
”The Ethiopian similarity with the Yemeni detected throughout the genome could be explained as an Ethiopian contribution to the Yemeni gene pool, consistent with that observed with mtDNA.”[24]
A 2015 study by Dobon et al. identified an ancestral autosomal component of West Eurasian origin that is common to many modern Afroasiatic-speaking populations in Northeast Africa. Known as the ‘’Coptic’’ component, it peaks among Egyptian Copts who settled in Sudan over the past two centuries. The Coptic component evolved out of a main Northeast African and Middle Eastern ancestral component that is shared by other Egyptians and also found at high frequencies among Cushites (≈40–60%), with the remaining 40–60% of the ancestry of Cushites being of sub-Saharan East African origin. However, these percentages are not certain since Figure 3 of the study by Dobon et al. only features these percentages in the k=3 model.[25] According to Dobon et al. (2015), the results were not conclusive and states that there needs to be more studies on the East African samples. Dobon et al. states:
”Our main results add new and interesting features to the North East African genetic complexity, with new populations that define a genetic component in southern Nilo-Saharan speakers that cannot be related to a North-African or other sub-Saharan components. These populations should be included in further population genetics and epidemiological studies to have a representative sample of the genetic diversity of the region of East Africa.”[25]
The scientists involved in the study suggest that this points to a common origin for the general population of Egypt.[25] They also associate the Coptic component with Ancient Egyptian ancestry, without the later Arabian influence that is present among other Egyptians.[25] The Coptic component is roughly equivalent with the Ethio-Somali component.[22]
Y DNA
changeA Y-chromosome study by Wood et al. (2005) tested various populations in Africa for paternal lineages, including 26 Maasai and 9 Luo from Kenya, and 9 Alur from the Democratic Republic of Congo. The signature Nilotic paternal marker Haplogroup A3b2 was observed in 27% of the Maasai, 22% of the Alur, and 11% of the Luo.[26]
According to Gomes et al. (2010),[27] Haplogroup B is another characteristically Nilotic paternal marker. It was found in 22% of Wood et al.’s Luo samples, 8% of studied Maasai and 50% of studied Nuer.[26] The E1b1b haplogroup has been observed at overall frequencies of around 11% among Nilo-Saharan-speaking groups in the Great Lakes area,[28] with this influence concentrated among the Maasai (50%).[26] This is indicative of substantial historic gene flow from Cushitic-speaking males into these Nilo-Saharan-speaking populations.[28] In addition, 67% of the Alur samples possessed the E2 haplogroup.[26]
A study by Hassan et al. (2008) analysed the Y-DNA of populations in the Sudan region, with various local Nilotic groups included for comparison. The researchers found the signature Nilotic A and B clades to be the most common paternal lineages amongst the Nilo-Saharan speakers, except those inhabiting western Sudan. There, a prominent North African influence was noted. Haplogroup A was observed amongst 62% of Dinka, 53.3% of Shilluk, 46.4% of Nuba, 33.3% of Nuer, 31.3% of Fur and 18.8% of Masalit. Haplogroup B was found in 50% of Nuer, 26.7% of Shilluk, 23% of Dinka, 14.3% of Nuba, 3.1% of Fur and 3.1% of Masalit. The E1b1b clade was also observed in 71.9% of the Masalit, 59.4% of the Fur, 39.3% of the Nuba, 20% of the Shilluk, 16.7% of the Nuer, and 15% of the Dinka.[29] Hassan et al. attributed the atypically high frequencies of the haplogroup in the Masalit to either a recent population bottleneck, which likely altered the community’s original haplogroup diversity, or to geographical proximity to E1b1b’s place of origin in North Africa. The researchers suggest that the clade “might have been brought to Sudan […] after the progressive desertification of the Sahara around 6,000–8,000 years ago”.[29] Henn et al. (2008) similarly observed Afro-Asiatic influence in the Nilotic Datog of northern Tanzania, 43% of whom carried the M293 sub-clade of E1b1b.[30]
mtDNA
changeUnlike the paternal DNA of Nilotes, the maternal lineages of Nilotes in general show low-to-negligible amounts of Afro-Asiatic and other extraneous influences. An mtDNA study by Castri et al. (2008) examined the maternal ancestry of various Nilotic populations in Kenya, with Turkana, Samburu, Maasai and Luo individuals sampled. The mtDNA of almost all of the tested Nilotes belonged to various Sub-Saharan macro-haplogroup L sub-clades, including L0, L2, L3, L4 and L5. Low levels of maternal gene flow from North Africa and the Horn of Africa were observed in a few groups, mainly via the presence of mtDNA haplogroup M and haplogroup I lineages in about 12.5% of the Maasai and 7% of the Samburu samples, respectively.[31]
Autosomal DNA
changeThe autosomal DNA of Nilotic peoples has been examined in a comprehensive study by Tishkoff et al. (2009) on the genetic clusters of various populations in Africa. According to the researchers, Nilotes generally form their own African genetic cluster. The authors also found that certain Nilotic populations in the eastern Great Lakes region, such as the Maasai, showed some additional Afro-Asiatic affinities due to repeated assimilation of Cushitic-speaking peoples over the past 5000 or so years.[32]
Admixture analysis
changeTishkoff ‘’et al’’ in 2009 published the largest study done to characterise genetic variation and relationships among populations in Africa. They examined 121 African populations, 4 African American populations and 60 non-African populations. Their results indicated a high degree of mixed ancestry reflecting migration events. In East Africa, all population groups examined had elements of Nilotic, Cushitic and Bantu ancestry amongst others to varying degrees. They also found that by and large, genetic clusters were consistent with linguistic classification with notable exceptions including the Luo of Kenya. Despite being Nilo-Saharan speakers, the Luo cluster with the Niger-Kordofanian speaking populations that surround them. They suggest that this indicates a high degree of admixture occurred during the southward migration of Southern Luo. Kalenjin groups and Maasai groups were found to have less Bantu ancestry but significant Cushitic ancestry.[32]
David Reich, a geneticist known for his studies on ancient DNA carried out a study which found that mutation frequencies in Luo people were much more similar to those of the surrounding Bantu speakers. They suggested that Luo speakers in East Africa may not have always been socially disadvantaged as they migrated into territories already inhabited by Bantu speakers. This is in keeping with oral history which affirms that large groups of Bantu speakers adopted Luo language, culture and customs that were dominant at the time.[33]
References
change- ↑ FALLON, PAUL D. "Language Development in Eritrea." Sustaining Linguistic Diversity: Endangered and Minority Languages and Language Varieties (2008): 145.
- ↑ University of Michigan (1987). Canadian Journal of African Studies: Le Journal Canadien Des Études Africaines.
conservative Arab countries (partly at the behest of the United States) put pressure on Somalia to forestall the growth of a pan-Horn
- ↑ Savà, Graziano, and Mauro Tosco. "La mort des langues en domaine chamito-sémitique." Faits de Langues 27.1 (2006): 279-290.
- ↑ Kitchen, Kenneth A. "Red Sea Harbours, Hinterlands and Relationships in Preclassical Antiquity." BAR INTERNATIONAL SERIES 1661 (2007): 131.
- ↑ Mitchell, Peter; Lane, Paul (2013-07-04). The Oxford Handbook of African Archaeology. OUP Oxford. ISBN 978-0-19-162615-9.
- ↑ Klees, Frank; Kuper, Rudolph (1992-01-01). New light on the Northeast African past : current prehistoric research: Contributions to a symposium, Cologne 1990. Heinrich-Barth-Institut.
- ↑ Hepburn, H. Randall; Radloff, Sarah E. (2013-03-14). Honeybees of Africa. Springer Science & Business Media. ISBN 978-3-662-03604-4.
- ↑ Daniel, Kendie (1988). NORTHEAST AFRICA AND THE WORLD ECONOMIC ORDER. Michigan, US. pp. 69–82.
{{cite book}}: CS1 maint: location missing publisher (link) - ↑ Project MUSE. (2020). Northeast African Studies. Retrieved March 22, 2020. "This distinguished journal is devoted to the scholarly analysis of Ethiopia, Eritrea, Djibouti, Somalia, and Sudan, as well as the Nile Valley, the Red Sea, and the lands adjacent to both."
- ↑ Milkias, Paulos. "The Political Spectrum of Western Education in Ethiopia." Journal of African Studies 9.1 (1982): 22.
- ↑ Cousin, Tracey L. "Eritrean and Ethiopian Civil War". ICE Case Studies. Archived from the original on 11 September 2007. Retrieved 20 July 2020.
- ↑ Underhill, P. A.; Shen, P.; Lin, A. A.; Jin, L.; Passarino, G.; Yang, W. H.; Kauffman, E.; Bonné-Tamir, B.; Bertranpetit, J. (November 2000). "Y chromosome sequence variation and the history of human populations". Nature Genetics. 26 (3): 358–361. doi:10.1038/81685. ISSN 1061-4036. PMID 11062480. S2CID 12893406.
- ↑ Morling, Niels; Hernandez, Alexis; Børsting, Claus; Hallenberg, Charlotte; Sanchez, Juan J. (July 2005). "High frequencies of Y chromosome lineages characterized by E3b1, DYS19–11, DYS392–12 in Somali males". European Journal of Human Genetics. 13 (7): 856–866. doi:10.1038/sj.ejhg.5201390. ISSN 1476-5438. PMID 15756297.
- ↑ Cruciani, Fulvio; Santolamazza, Piero; Shen, Peidong; Macaulay, Vincent; Moral, Pedro; Olckers, Antonel; Modiano, David; Holmes, Susan; Destro-Bisol, Giovanni (May 2002). "A back migration from Asia to sub-Saharan Africa is supported by high-resolution analysis of human Y-chromosome haplotypes". American Journal of Human Genetics. 70 (5): 1197–1214. doi:10.1086/340257. ISSN 0002-9297. PMC 447595. PMID 11910562.
- ↑ Semino, Ornella; Santachiara-Benerecetti, A. Silvana; Falaschi, Francesco; Cavalli-Sforza, L. Luca; Underhill, Peter A. (January 2002). "Ethiopians and Khoisan share the deepest clades of the human Y-chromosome phylogeny". American Journal of Human Genetics. 70 (1): 265–268. doi:10.1086/338306. ISSN 0002-9297. PMC 384897. PMID 11719903.
- ↑ Moran, Colin N.; Scott, Robert A.; Adams, Susan M.; Warrington, Samantha J.; Jobling, Mark A.; Wilson, Richard H.; Goodwin, William H.; Georgiades, Evelina; Wolde, Bezabhe (2004-10-20). "Y chromosome haplogroups of elite Ethiopian endurance runners". Human Genetics. 115 (6): 492–497. doi:10.1007/s00439-004-1202-y. ISSN 0340-6717. PMID 15503146. S2CID 13960753.
- ↑ Shen, Peidong; Lavi, Tal; Kivisild, Toomas; Chou, Vivian; Sengun, Deniz; Gefel, Dov; Shpirer, Issac; Woolf, Eilon; Hillel, Jossi (2004). "Reconstruction of patrilineages and matrilineages of Samaritans and other Israeli populations from Y-Chromosome and mitochondrial DNA sequence Variation". Human Mutation. 24 (3): 248–260. doi:10.1002/humu.20077. ISSN 1059-7794. PMID 15300852. S2CID 1571356.
- ↑ Ehret, Christopher (2008), "Reconstructing Ancient Kinship in Africa", Early Human Kinship, Blackwell Publishing Ltd., pp. 200–231, doi:10.1002/9781444302714.ch12, ISBN 9781444302714
- ↑ Kivisild, Toomas; Reidla, Maere; Metspalu, Ene; Rosa, Alexandra; Brehm, Antonio; Pennarun, Erwan; Parik, Jüri; Geberhiwot, Tarekegn; Usanga, Esien (November 2004). "Ethiopian Mitochondrial DNA Heritage: Tracking Gene Flow Across and Around the Gate of Tears". American Journal of Human Genetics. 75 (5): 752–770. doi:10.1086/425161. ISSN 0002-9297. PMC 1182106. PMID 15457403.
- ↑ Mikkelsen, Martin; Fendt, Liane; Röck, Alexander W.; Zimmermann, Bettina; Rockenbauer, Eszter; Hansen, Anders J.; Parson, Walther; Morling, Niels (July 2012). "Forensic and phylogeographic characterisation of mtDNA lineages from Somalia". International Journal of Legal Medicine. 126 (4): 573–579. doi:10.1007/s00414-012-0694-6. ISSN 1437-1596. PMID 22527188. S2CID 22566302.
- ↑ Loredana, Castrì. Kenyan crossroads : migration and gene flow in six ethnic groups from eastern Africa. OCLC 756782632.[page needed]
- ↑ 22.0 22.1 22.2 22.3 Hodgson, JA; Mulligan, CJ; Al-Meeri, A; Raaum, RL (June 2014). "Early back-to-Africa migration into the Horn of Africa". PLOS Genetics. 10 (6): e1004393. doi:10.1371/journal.pgen.1004393. PMC 4055572. PMID 24921250.
- ↑ Lazaridis, Iosif; Nadel, Dani; Rollefson, Gary; Merrett, Deborah C.; Rohland, Nadin; et al. (25 July 2016). "Genomic insights into the origin of farming in the ancient Near East". Nature. 536 (7617): 419–424. Bibcode:2016Natur.536..419L. doi:10.1038/nature19310. PMC 5003663. PMID 27459054.
- ↑ 24.0 24.1 Pagani, Luca; Kivisild, Toomas; Tarekegn, Ayele; Ekong, Rosemary; Plaster, Chris; et al. (13 July 2012). "Ethiopian Genetic Diversity Reveals Linguistic Stratification and Complex Influences on the Ethiopian Gene Pool". The American Journal of Human Genetics. 91 (1): 83–96. doi:10.1016/j.ajhg.2012.05.015. PMC 3397267. PMID 22726845.
- ↑ 25.0 25.1 25.2 25.3 Dobon, B; Hassan, HY; Laayouni, H; Luisi, P; Ricaño-Ponce, I; et al. (28 May 2015). "The genetics of East African populations: a Nilo-Saharan component in the African genetic landscape". Scientific Reports. 5: 9996. Bibcode:2015NatSR...5.9996D. doi:10.1038/srep09996. PMC 4446898. PMID 26017457.
- ↑ 26.0 26.1 26.2 26.3 Wood et al. 2005, pp. 867–876.
- ↑ Gomes et al. 2010, pp. 603–13.
- ↑ 28.0 28.1 Cruciani et al. 2004, pp. 1014–1022.
- ↑ 29.0 29.1 Hassan et al. 2008, pp. 316–323.
- ↑ Henn et al. 2008, pp. 10693–10698.
- ↑ Castrì et al. 2008, pp. 189–92.
- ↑ 32.0 32.1 Tishkoff et al. 2009, pp. 1035–44.
- ↑ Reich 2018, pp. 215–216.